pset 06: Student's game night
On Tuesday nights, the lab's favorite pub is tended by an odd bartender. Nobody knows his real name. He calls himself Student, and he serves only Guinness. Student challenges customers to play a strange betting game of his own invention.
the game
In a back room, hidden from the customers, there is a 20 row x 21 column grid on the wall. The rows are called \(\sigma\), and they are marked 5, 10, 15, up to 100, starting from the bottom. The 21 columns are called \(\mu\), and they are labeled -100, -90, -80 and so on up to 100, starting from the left.
Student, a practiced dart player with the uncanny ability to throw a dart into a rectangular grid with a perfectly uniform distribution, says he starts each game by stepping into the back room and throwing a dart at the board. Thus, he randomly selects one of the squares with uniform probability, thus uniformly sampling from discrete values \(\mu\) and \(\sigma\).
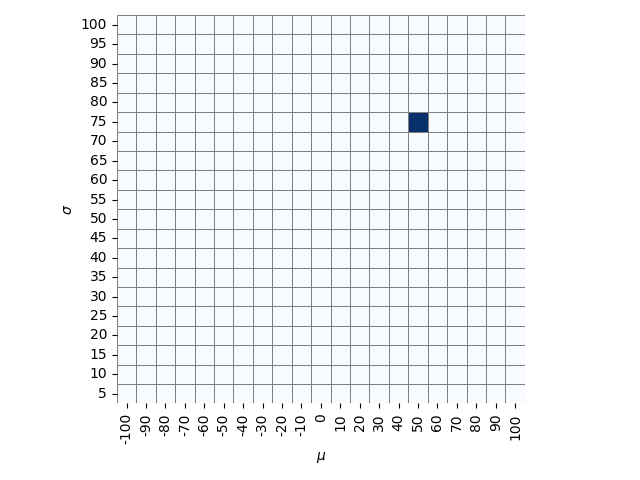 An example of the grid in Student's back room, where he's thrown a dart
that hit mu = 50, sigma = 75.
An example of the grid in Student's back room, where he's thrown a dart
that hit mu = 50, sigma = 75.
Student, unsurprisingly, is also an eccentric inventor. He explains that once the dart selects \(\mu\) and \(\sigma\), an elaborate hidden machine is activated in the ceiling. (You think you hear him say the word quincunx, but maybe it was only a sneeze.) The machine drops an object such that the object lands along a line on the bar, with a position that is Gaussian-distributed with mean \(\mu\) and standard deviation \(\sigma\). The line on the bar is carefully marked off as a real number line, in very fine increments, so customers can measure object positions precisely. Strangely, though you hadn't noticed before, now you realize that the bar appears to be infinitely long. Student says the positions of these \(n\) objects will be called \(x_i\) for \(i = 1..n\).
Student explains that his machine used to drop ping pong balls, but they rolled away. He found it works better with objects that are soft, so they stay where they land, and can land on top of each other, so now the machine uses tea bags instead. Everyone calls it Student's tea distribution machine.
In summary (in case you didn't follow all the unnecessary nonsense around the problem): Student uniformly samples a mean \(\mu\) from 21 possible choices, and a standard deviation \(\sigma\) from 20 possible choices. From a normal distribution parameterized by \(\mu\) and \(\sigma\), he samples \(n\) numbers \(x_i\), \(i=1..n\) that you observe. Depending on the game, Student may or may not give you additional information about \(\mu\) and \(\sigma\).
Student reminds you it's a rookie mistake to confuse the observed data \(x_1..x_n\) with the unknown parameters of the machine \(\mu\) and \(\sigma\). He says to remember that the hidden grid has only 20x21 discretized choices of \(\sigma\) and \(\mu\), but the tea bags can land anywhere along the line (i.e. the observations are continuously distributed, but the parameters are discretized).
The game is to guess where what column the hidden dart is in: i.e., what the unknown mean \(\mu\) is.
the betting
Customers are allowed to bet after each tea bag is dropped (that is, after each observation). There's a complicated display in the pub that keeps track of the observations and the betting. Betting odds are updated instantly at each round, based on the observations that have been seen so far. There are two different versions of the game:
-
the beginner's game: During a beginner round, Student discloses what row the dart is in (what \(\sigma\) is), but not the column. So the game is to deduce \(\mu\), given known \(\sigma\) and the observed data \(x_1..x_n\).
-
the advanced game: In an advanced round, you have to deduce the column \(\mu\) just from the observed data \(x_1..x_n\); now \(\sigma\) is unknown.
One thing in particular catches your attention. The posted rules are explicit about how the pub estimates fair odds of the game mathematically from the current observed samples \(x_1..x_n\), as follows:
-
the sample mean \(\bar{x} = \frac{1}{n} \sum_{i=1}^{n} x_i\) is calculated;
-
the sample standard deviation \(s = \sqrt{ \frac{ \sum_{i=1}^{n} (\bar{x} - x_i)^2 } {n-1} }\) is calculated;
-
in the beginner's game, with \(\sigma\) known, the unknown true location of \(\mu\) is assumed to be distributed proportional to a normal distribution with mean \(\bar{x}\) and standard deviation \(\frac{\sigma}{\sqrt{n}}\). (You recognize the familiar equation for the standard error of the mean, when the parametric \(\sigma\) is known.)
-
in the advanced game, the unknown true location of \(\mu\) is assumed to be distributed proportional to a normal distribution with mean \(\bar{x}\) and standard deviation \(\frac{s}{\sqrt{n}}\): i.e. using the observed sample standard deviation as an estimate of \(\sigma\), rather than a known parameter \(\sigma\). Again you recognize a familiar equation for the standard error of the mean. (See the notes, below, for why this says "proportional to" a normal distribution; short answer: \(\mu\) isn't continuous here, it comes in 21 discrete values on the hidden grid.)
Student may be brilliant and all, but you realize there's something not right about the way Student is calculating the odds for the advanced game, especially when the sample size \(n\) is small. If you can do a better job of inferring the hidden column position of the dart -- the unknown \(\mu\) -- you can use that information to your betting advantage. Now that you know a few things about Bayesian inference, you set about to calculate the posterior distributions \(P( \mu \mid x_1..x_n, \sigma)\) and \(P( \mu \mid x_1..x_n)\) directly, without resorting to traditional summary statistics like the sample mean and sample standard deviation. Using these inferred distributions, you'll know the true odds, and be able to take advantage when the pub has miscalculated the odds.
You can download
a script that implements Student's game. To use
it, do ./student-game.py <n>, where <n> is the number of
observed tea bag positions. For example:
% ./student-game.py 3
generates 3 samples, along with all the other information about the game, including the hidden correct answers (so you can check how well your inference works, as \(n\) varies.)
An example to use in your analysis below:
x = np.array([ 11.50, -2.32, 9.18]) # n=3 observations
true_sigma = 60. # Student also tells you this in beginner's game
true_mu = -20. # the unknown column position, mu
This is an interesting example because all three samples just happened to come out to the right of the true \(\mu\), by chance, and fairly tightly grouped. (It's a real example that came up in testing this exercise.)
1. the beginner's game
Write a script that:
-
takes \(n\) observations \(x_1..x_n\) and one of the 20 possible values of \(\sigma\) (i.e. a known row, specified by Student) as input.
-
calculates the posterior probability \(P( \mu \mid x_1..x_n, \sigma)\) for each of the 21 possible values of \(\mu\) on that grid row. Remember that the prior \(P(\mu)\) is uniform.
-
plots that distribution on a semilog scale (so you can see differences in the small-probability tail more easily), using the
semilogyplot ofmatplotlibfor example. -
and plots the pub's calculated probability distribution on the same semilog plot, so you can compare.
You can use this script, which implements Student's game, to generate data (and \(\sigma\)) to try your analysis out, for varying numbers of samples, especially small \(n\) (3-6).
Have your script show the plots for the x = [ 11.50, -2.32, 9.18],
true_sigma = 60. example.
2. the advanced game
Now write a second script that:
-
just takes \(n\) observations \(x_1..x_n\) (now \(\sigma\) is unknown).
-
calculates the posterior probability \(P( \mu, \sigma \mid x_1..x_n)\) for each of the 420 (20x21) possible values of \(\sigma,\mu\) on Student's grid.
-
plots that 20x21 posterior distribution as a heat map
-
marginalizes (sum over the rows) to obtain \(P(\mu \mid x_1..x_n)\):
-
plots that marginal distribution on a semilog scale;
-
and plots the pub's calculated probability distribution, so you can compare.
Again you can use the Student's game script to generate data (and \(\sigma\)) to try your analysis out, for varying numbers of samples, especially small \(n\) (3-6).
Have your script show the plots for the X = [ 11.50, -2.32, 9.18] example.
3. where's the advantage?
Is the pub calculating its odds correctly? Where do you see an advantage?
(i.e. why is there a difference, and how could you take advantage of this difference in betting?)
allowed modules
NumPy, Matplotlib, Pandas, Seaborn, and SciPy's stats and special modules.
You'll probably want stats.norm.logpdf() for the Gaussian log PDF,
and special.logsumexp() for summing probabilities that you're
storing as log probabilities.
I like Seaborn's sns.heatmap() for heat maps.
You probably don't need Pandas.
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as stats
import scipy.special as special
import seaborn as sns
import pandas as pd # optional
%matplotlib inline
turning in your work
Submit your Jupyter notebook page (a .ipynb file) to
the course Canvas page under the Assignments
tab.
Name your
file <LastName><FirstName>_<psetnumber>.ipynb; for example,
mine would be EddySean_06.ipynb.
We grade your psets as if they're lab reports. The quality of your text explanations of what you're doing (and why) are just as important as getting your code to work. We want to see you discuss both your analysis code and your biological interpretations.
Don't change the names of any of the input files you download, because the TFs (who are grading your notebook files) have those files in their working directory. Your notebook won't run if you change the input file names.
Remember to include a watermark command at the end of your notebook,
so we know what module versions you used, in case we have to
reproduce some bug exactly. For example,
%load_ext watermark
%watermark -v -m -p numpy,matplotlib,seaborn,pandas,jupyter
the point of this baroque exercise (there is one)
Student's t distribution arises when we're trying to estimate the mean of a normal distribution from observed data, and the variance is unknown. Bayesian inference tells us that the posterior distribution of \(P(\mu \mid x_1..x_n)\) is:
where we have to integrate (marginalize) over the unknown and uncertain \(\sigma\) parameter.
The Student's tea distribution machine problem discretizes the choices for \(\mu\) and \(\sigma\) to create a toy problem where we can sum instead of integrate, and it explicitly makes their prior distributions uniform so the posterior distribution over \(\mu\) is well defined. This suffices to demonstrate the general properties and shape of the real Student's t distribution, with its fat tails relative to the normal distribution.
The pub should be using the t-distribution to calculate its odds:
def probdist_t(X, mu_values):
"""
Given an ndarray X_1..X_n,
and a list of the mu values in each column;
return a list of the inferred P(mu | X) for each column,
according to Student's t distribution with N-1 degrees of freedom.
"""
N = len(X)
xbar = np.mean(X)
s = np.std(X, ddof=1)
t = [ (xbar - mu) / (s / np.sqrt(N)) for mu in mu_values ] # t statistic, given sample mean, sample stddev, and N
# t = [ stats.ttest_1samp(X, mu)[0] for mu in mu_values ] # ... (equivalently, python can calculate t statistic for you)
Pr = [ stats.t.pdf(val, N-1) for val in t ]
Z = sum(Pr)
Pr = [ p / Z for p in Pr ]
return Pr
You can add this line to your plot for the advanced game, to see how it roughly matches your Bayesian calculation.
With a small number of samples, your uncertainty of your estimate for the unknown \(\sigma\) becomes important, and making a point estimate for it isn't a good idea. The correct thing to do is to marginalize over the uncertain and unknown hidden parameter \(\sigma\).
Although Student derived it analytically (which was quite a feat!) and not as a Bayesian calculation, the exercise illustrates how Student's t distribution can be seen to arise as the marginal posterior probability for \(\mu\), given observed data, marginalized over unknown \(\sigma\) with a uniform prior.
hints
-
The script used to produce the figure showing Student's dart grid is here. It uses a Seaborn heat map, so you can use it as an example of plotting heat maps.
-
The reason to say "proportional to" a normal distribution is that there are only 21 discrete values of \(\mu\) possible in the game. The rule board (and the Student's game script) include a helpful Python code snippet, the implementation of the pub's rules for calculating fair odds:
def probdist_beginner(X, sigma, mu_values):
"""
Given an ndarray X_1..X_n, and a known sigma;
and a list of the mu values in each column;
return a list of the pub's inferred P(mu | X,sigma) for each column.
"""
xbar = np.mean(X)
N = len(X)
Pr = [ stats.norm.pdf(x, loc=xbar, scale= sigma / math.sqrt(N)) for x in mu_values ] # proportional to std error of the mean
Z = sum(Pr) # normalization constant
Pr = [ p / Z for p in Pr ] # normalization to a discrete probability distribution
return Pr
def probdist_advanced(X, mu_values):
"""
Given an ndarray X_1..X_n,
and a list of the mu values in each column;
return a list of the pub's inferred P(mu | X) for each column.
"""
xbar = np.mean(X)
s = np.std(X, ddof=1) # note that numpy.std() by default calculates a population std dev; to get sample std. dev., set ddof=1
N = len(X)
Pr = [ stats.norm.pdf(x, loc=xbar, scale= s / math.sqrt(N)) for x in mu_values ] # proportional to std error of the mean
Z = sum(Pr) # normalization constant
Pr = [ p / Z for p in Pr ] # normalization to a discrete probability distribution
return Pr
- To be precise about the gambling part: in statistics, "odds" means \(\frac{p}{1-p}\) for some outcome that occurs with probability \(p\): the ratio of the probability that the outcome occurs versus doesn't occur. In gambling, odds are typically expressed in terms of the payout if the outcome occurs. For example, 5:1 odds means if you bet a dollar and you win, you win five dollars (in addition to keeping your one dollar bet). When the house offers fair odds, that means they set the payout so that the expected value is 0, which means setting the payout to 1/odds, i.e. to \(\frac{1-p}{p}\) : 1. For example, if the probability that Moriarty gets his PhD is 10%, and you were going to bet me on it, 9:1 would be fair odds for me to offer you.
In the pset, this gambling jargon is not the point of the exercise. You only need to focus on calculating correct posterior probabilities, and on whether Student is over- or under-estimating them. The idea is that if Student miscalculates the probability, he will offer you "fair odds" that are wrong (where your expected winnings aren't zero); if you can identify situations where your expected winnings are positive (where Student has underestimated the probability), you can exploit that.